Tree simulation is a crucial instrument for exploring the processes that will have generated biodiversity. A typical instrument for simulating trees is the start–dying simulation or equals-rates Markov mannequin (ERMM) that randomly removes and provides tips about a tree Many publications have independently explored alterations of the ERMM, similar to modifying the charges of speciation for different suggestions within the tree
All such research, nonetheless, have developed their very own software program instruments for tree simulation. The TreeMan object is definitely modified using the addTip() and rmTip() capabilities and its pace of processing and implementation makes TreeMan a super software program instrument for tree simulation. Furthermore, as half of the R setting a consumer has entry to the wide selection of eco-evolutionary R packages already out there to broaden on these earlier tree simulations.
To exhibit this performance, we produced an R script for simulating a tree using an Evolutionary Distinctness Biased Markov Model (EDBMM with fewer than 40 strains of R code
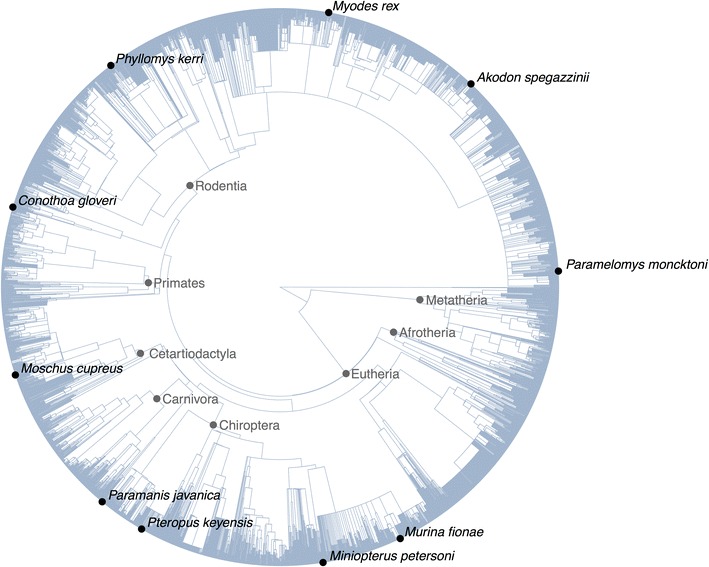
The simulation randomly provides and removes suggestions however with a bias in the direction of evolutionary distinctness: extra evolutionarily distinct suggestions have a decrease fee of speciation and extinction The equal script for working a vectorised EDBMM with a phylo object takes 188 strains (see ‘edbmm-with-phylo.R’ in Additional
A typical want in organic evaluation is the detection of phylogenetic sign. Such analyses are sometimes executed using model-based statistical analyses for which APE has been primarily designed. A model-based method, nonetheless, is just not all the time relevant to each query associated to phylogenetic sign. One such query is whether or not the adjustments in species between habitat varieties are as a consequence of phylogenetic sign in species’ good points and losses, e.g.
consequently of human-caused habitat loss]. A helpful metric for calculating such a distinction is the UniFrac measure , which measures the distinctive and shared fractions of branches represented between communities.
The TreeMan object is nicely suited to calculating this metric as all nodes in a tree have IDs; even when nodes are added or eliminated, IDs are fixed. To exhibit this we randomly generated neighborhood information with different intensities of overlap. We then ran permutation assessments that detect whether or not there was vital phylogenetic turnover between these communities using treeman’s calcOvrlp
Code snippet for calculating overlap between two different communities using treeman and ExtraTreeTool: generate random communities using parameters of neighborhood overlap for a random tree, plot as neighborhood trees , convert trees from phylo to TreeMan using ExtraTreeToolas(), generate null communities to check whether or not the 2 communities have vital overlap